Popular
Expert interviews
ToxicologyIn silicoPolicy
Tox 21: A New Way to Evaluate Chemical Safety and Assess Risk
Tox21 is a US federal research collaboration focused on driving the evolution of Toxicology in the 21st Century by developing methods to rapidly and efficiently evaluate the safety of commercial chemicals, pesticides, food additives/contaminants, and medical products. The goals of Tox21 are to (1) identify mechanisms of chemically-induced biological activity; (2) prioritize chemicals for more extensive testing; and (3) develop more relevant and predictive models of in vivo toxicological responses.

Innovation examples
HealthToxicology
Zebrafish in toxicity testing
Zebrafish are increasingly recognised as a useful model for toxicity testing of chemical substances. Testing strategies are becoming more based on mechanisms of toxicity structured in adverse outcome pathways describing the chain of events leading to toxicity or disease. Using a battery of dedicated in vitro and in silico assays, insight can be gained in how exposure leads to disease. For certain diseases it is known that toxicity relies on the interaction between different organs and cell types, which requires research on whole organisms in addition to simple in vitro models. The zebrafish is considered a valuable whole organism model in a mechanism-based testing strategy. At RIVM, the zebrafish embryo model is used for testing the effect of chemical substances on several adverse outcomes and diseases.
For more information see: https://ehp.niehs.nih.gov/doi/10.1289/EHP9888; https://doi.org/10.3390/ijerph18136717; www.linkedin.com/in/harm-heusinkveld

Innovation examples
ToxicologyIn vitroOrgan-on-Chip
Cartilage-on-a-chip for studying joint degenerative diseases
Carlo Alberto Paggi is currently a PhD candidate at the University of Twente in the research group of Prof. Marcel Karperien and Prof. Séverine Le Gac. Karperien’s lab focus on the biological aspects of osteoarthritic research while Le Gac’s specialize in organ-on-chip development. The project of Carlo Alberto is developing a joint-on-chip platform to create a reliable in vitro model to study disease progression in osteo- or rheumatoid arthritis. The model combines different organ-on-chips aimed at replicating each a tissue around the joint such as cartilage, bone and ligaments. This new technology focuses on better reproducing human models and at substituting the use of animal models for drug research. If you want to know something more about the project and the groups, you can follow the link in the video.
Carlo Paggi was nominated for the Hugo van Poelgeest prize for his research on a cartilage-on-a-chip model to study joint degenerative diseases
Karperien’s lab of Developmental Bioengineering: https://www.utwente.nl/en/tnw/dbe/
Le Gac’s lab of Applied Microfluidics for BioEngineering Research: http://www.severinelegac.com/
Linkedin: https://www.linkedin.com/in/carlo-alberto-paggi-76500b135/
Meetings & conferences
HealthIn vitroAdvanced
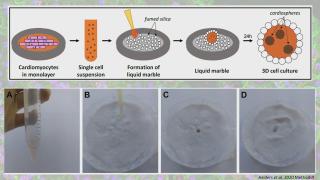
Liquid marbles for cardiac organoids development
Advances in three-dimensional (3D) culture techniques have shown several advantages over 2D cultures, especially by more accurately mimicking the in vivo environment. This has led to improved reproducibility and reliability of experimental results, which are important criteria in disease modelling and toxicity testing. Induced pluripotent stem cells (iPSC) provide an unlimited source for the derivation of all cell types of the adult body, including cardiomyocytes. To improve the current culture methods for multicellular cardiac spheroids, such as the hanging drop method, we explored the use of hydrophobic powders. Fumed silica nanoparticles can be used to encapsulate liquid drops, which could serve as a microenvironment for cell cultures. This microbioreactor stimulates cell coalescence and 3D aggregation while providing optimal gas exchange between the interior and the surrounding environment. Moreover, the properties of liquid marble microbioreactors render them ideal for co-culture experiments. This liquid marble technique has been previously explored and optimized for other cell types. Here we describe a protocol that allows for the derivation of functional cardiac mini organoids, consisting of co-cultured cardiomyocytes and cardiac fibroblasts. These cardiospheres can be valuable for modelling cardiac diseases in vitro and assessing cell interactions to decipher disease mechanisms.
Lab website: https://www.medicalcellbiologylab.com/
Contact: https://www.researchgate.net/profile/Jeffrey-Aalders
RE-place database: https://www.re-place.be/method/liquid-marbles-cost-effective-platform-generate-cardiospheres-co-cultured-cardiomyocytes-and

Innovation examples
HealthIn vitroOrgan-on-Chip
An iPSC-derived blood-brain barrier to model neurodegeneration
The blood-brain barrier is a layer of cells that protects our brain from harmful compounds. However, due to this tight barrier, many drugs to treat neurological diseases cannot enter the brain either.
There are currently no good models to test these types of drugs. Henrique Nogueira Pinto is a PhD candidate at the Vrije Universiteit in Amsterdam. He is developing a blood-brain barrier model coupled to mini-brains. With this model, he aims to more reliably test how drugs can be transported over the blood-brain barrier and what their effect on the brain is.
Click on the info button for the full version of the video. Click here (https://fluidsbarrierscns.biomedcentral.com/articles/10.1186/s12987-022-00316-0#Sec3) for a review of the current status of in vitro models for the blood-brain barrier.

Innovation examples
ToxicologyIn vitroOrgan-on-Chip
Human pluripotent stem cell derived cardiomyocytes for disease modelling and drug discovery
Berend van Meer did his PhD research in the research group of prof. Christine Mummery at the department of Anatomy and Embryology of the Leiden University Medical Center. In this group, human pluripotent stem cell derived (Organ-on-Chip) models are being developed, mostly cardiovascular models. The work of Berend aimed to understand how well these stem cell based cardiac models can predict the effect of (well-known) drug therapies in patients. Importantly, the outcomes of the experiments were compared to very similar measurements in rabbit heart muscle cells. And while animal models predicted less than 70% correctly, the human stem cell based models predicted almost 80% of the expected effects correctly. The research contributes to understanding the relevance of stem cell based models and strengthens the confidence regulators and pharmaceutical companies have in such models as animal alternatives in the drug development pipeline.
Berend van Meer has won the Hugo van Poelgeest prize 2020 for his research on human pluripotent stem cell derived cardiomyocytes for disease modelling and drug discovery.
Christine Mummery's lab on Heart on Chip, Disease modeling and toxicity: https://www.lumc.nl/org/anatomie-embryologie/research/902040935402533/
Meetings & conferences
HealthIn vitroAdvanced
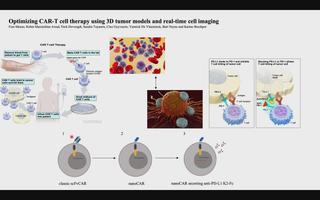
3D tumor models for CAR-T-cell therapy optimization
Chimeric antigen receptor (CAR) T-cell therapy accounts for one of the most promising therapeutic advances in cancer immunotherapy. In this form of adoptive cell transfer, T-cells of a patient are engineered to express so-called ‘CARs’, in which the antigen-recognition capacity of antibodies is combined with T-cell activating domains. So far, CAR-T-cell therapy obtained its most impressive results in hematological malignancies resulting in the approval of five CAR-T cell products by the FDA for hematologic indications. However, CAR-T-cell therapy has not mirrored its success in solid tumors. The poor efficacy of CAR-T-cell therapy in solid tumors has, in part, been attributed to the lack of understanding in how CAR-T-cells function in a solid tumor microenvironment. Classical validation methods rely on the use of specificity and functionality assays in 2D models against adherent target cells or target cells in suspension. Yet, by using these models, observations made in vitro may differ greatly to an in vivo situation where tumors are engrafted in 3D structures. We developed a more relevant and translational 3D tumor model using eGFP+ target cells. This allows us to couple 3D tumor cell killing by CAR-T-cells to live-cell imaging, providing an efficient quantification of target cell death. As proof- of-concept, we used a 3D model of eGFP+ glioblastoma cells and CAR-T-cells targeting a pan-cancer antigen. This 3D glioblastoma model allowed us to show that classical scFv-based CAR-T-cell therapy of glioblastoma cells can be improved by nanoCAR-T-cells. Furthermore, combining nanoCAR-T-cell therapy with a genetic approach of nanobody-based anti-PD-L1 immune checkpoint blockade further increased the cytotoxicity of the nanoCAR-T-cell therapy.

Innovation examples
HealthInnovationIn silico
Using data and computational modelling in biomedical research
Bioinformatics and systems biology hold great promise to translate the wealth of biological data into meaningful knowledge about human health and disease. The group of Bas Teusink helps biologists to deal with high throughput data, for example metabolomics (how cell metabolism works) and proteomics (how protein networks work) from patient material or cell cultures. This can help to better understand disease mechanisms and aid drug targeting or personalised medicine. In the future, combining data from different models (in vitro, in vivo and human data) could become a digital model of humans, or a “ digital twin”.
Click on the link in the video to watch more or read the interview with Bas (and Jaap Heringa) he[https://vu.nl/en/research/more-about/using-data-and-computational-modelling-in-biomedical-research]re.

Innovation examples
In vitroOrgan-on-Chip
Unified organoid system for modeling heart and kidney interaction on-a-chip
Beatrice Gabbin is a PhD candidate at the Anatomy and Embryology Department of the Leiden University Medical Center. Her project is shared with the Nephrology Department and focusses on the study of the cardiorenal axis in vitro. Both heart and kidneys have vital functions in the human body and reciprocally influence each other’s behavior: pathological changes in one can damage the other. There are already multiple independent in vitro (human) models of heart and kidney, but none have so far captured their dynamic crosstalk. The aim of the project is therefore to develop a microfluidic system which can be used to study heart and kidney interaction in vitro. For this purpose, cardiac microtissues and kidney organoids derived from human induced pluripotent stem cells are generated and loaded onto a 3D perfusion chip for their dynamic co-culture. This system enables the study the cardiac and kidney interaction with a high level of control. The validation of a unified organoid system will enable the investigation of diseases involving the two organs and their potential treatments. Read more via the link in the video and https://doi.org/10.1016/j.mtbio.2023.100818.

TPI.tv videos
InnovationPolicyBeginner
TPI.tv: improving science through animal-free innovations and research
Introducing TPI.tv : a video platform by experts striving to improve science through animal-free innovations and research.

Innovation examples
HealthToxicologyIn vitroOrgan-on-Chip
GUTS BV - small intestine-on-a-chip and advanced computational analysis for compound and protein screening
GUTS BV is a contract research organization offering its 3-dimensional state-of-the-art small intestinal in vitro model in combination with custom computational analysis approaches. The small intestinal model was developed during Dr. Paul Jochems PhD research at Utrecht University in the group of Prof. Roos Masereeuw. In comparison to the current gold standard (Transwell model), they show improvement in cell differentiation (all major specialized cell types present), physiological structure (3D tube- and villi-like structures) and a functional epithelial barrier. After acquiring experimental data from this model computational analysis approaches are used to score and compare measured compounds for all tested biological parameters at once. The combined effort of improved in vitro modelling and data analysis is believed to result in an enhanced preclinical predictability.
GUTS BV was nominated for the Venture Challenge 2021 for their development of an intestinal model combined with advanced computational analysis for protein and chemical compound screening.
Research papers:
https://www.sciencedirect.com/science/article/pii/S0887233318307811
https://www.mdpi.com/2072-6643/12/9/2782/htm
https://www.nature.com/articles/s41538-020-00082-z
LinkedIn: https://www.linkedin.com/company/71016128/

Innovation examples
HealthIn vitro
Immortalized human cells to model atrial fibrillation in vitro
Niels Harlaar is a PhD Candidate at the Laboratory of Experimental Cardiology at the Leiden University Medical Center. Here, under the supervison of prof. dr. D.A. Pijnappels and dr. A.A.F. de Vries, he focusses on the conditional immortalization of human atrial cardiomyocytes for (among many other applications) in vitro modelling of atrial fibrillation. He has successfully generated, characterized and applied this technique of these conditionally immortalized human atrial myocyte lines to model atrial fibrillation in vitro.
Niels won the Hugo van Poelgeest prize 2022 for excellent research to replace animal testing.
Click here (https://hartlongcentrum.nl/research/laboratory-of-experimental-cardiology/) for more information on the Laboratory of Experimental Cardiology.